Table Of Contents:
1. TCGA (The Cancer Genome Atlas)
2. CLEVR (Compositional Language and Elementary Visual Reasoning)
3. WIT (Wikipedia-based Image Text Dataset)
4. LAION-5B
5. MS COCO (Microsoft Common Objects in Context)
6. VQA (Visual Question Answering)
7. MIMIC-IV
8. UK Biobank
9. Human Cell Atlas (HCA)
10. TCIA (The Cancer Imaging Archive)
11. MLOmics
12. Kinetics-700
13. Flickr30k
14. ENCODE
15. Allen Brain Atlas
Multimodal datasets combine multiple data types like text, images, audio, video, genomics, and clinical data to create richer AI training resources. These datasets are crucial for multimodal deep learning, which requires integrating multiple data sources to enhance performance in tasks such as image captioning, medical diagnostics, scientific discovery, autonomous systems, and cross-modal analysis.
Professionals working with multimodal data need access to high-quality datasets that provide the comprehensive information required for training sophisticated AI models. The datasets featured in this guide represent the most valuable resources available for advancing multimodal machine learning research and applications across computer vision, natural language processing, and life sciences.
What is a multimodal dataset?
A multimodal dataset contains information from multiple data modalities such as text, images, audio, video, genomics, clinical records, and sensor data. Multimodal datasets bring different modalities together, helping researchers build AI models that can understand complex environments and reason across multiple information sources.
Multimodal datasets are the digital equivalent of our senses. Just as we use sight, sound, and touch to interpret the world, these datasets combine various data formats to offer a richer understanding of content and enable more sophisticated analysis.
Multimodal datasets enable AI systems to process information more holistically by combining complementary data types. This approach mirrors human cognition, where we naturally integrate information from multiple sources to understand our environment.
Dataset name | Dataset size | License | Authors | Ideal for |
TCGA | 20K+ samples, 2.5 PB | Open access | NCI/NHGRI | Cancer genomics |
CLEVR | 1M images + questions | BSD | Johnson et al. | Visual reasoning |
WIT | 37.6M image-text pairs | CC BY-SA 3.0 | Srinivasan et al. | Multilingual learning |
LAION-5B | 5.85B image-text pairs | Various | LAION collective | Large-scale training |
MS COCO | 330K images | CC BY 4.0 | Microsoft | Object detection |
VQA | 265K images + questions | CC BY 4.0 | Antol et al. | Question answering |
MIMIC-IV | 400K+ admissions | PhysioNet License | Johnson et al. | Healthcare AI |
UK Biobank | 500K+ participants | Approved access | UK Biobank | Population health |
Human Cell Atlas | Millions of cells | Open access | HCA Consortium | Single-cell biology |
TCIA | 30M+ images | Various | Clark et al. | Medical imaging |
MLOmics | 8,314 cancer samples | Open access | Chen et al. | Cancer ML |
Kinetics-700 | 650K video clips | YouTube ToS | Carreira et al. | Action recognition |
Flickr30k | 31K images + captions | Academic use | Young et al. | Image-text retrieval |
ENCODE | 1,600+ experiments | Open access | ENCODE Consortium | Gene regulation |
Allen Brain Atlas | Multi-brain regions | Open access | Allen Institute | Neuroscience |
To analyze multimodal datasets, specialized platforms and tools are required that can handle the complexity of multiple data types simultaneously. TileDB provides a unified platform for storing, organizing, and analyzing multimodal data using multi-dimensional arrays that can efficiently represent complex data structures.
1. TCGA (The Cancer Genome Atlas)
TCGA is a landmark cancer genomics program that molecularly characterized over 20,000 primary cancers and matched normal samples spanning 33 cancer types. This joint effort between NCI and the National Human Genome Research Institute generated over 2.5 petabytes of genomic, epigenomic, transcriptomic, and proteomic data.
TCGA combines multiple omics modalities including whole genome sequencing, RNA sequencing, DNA methylation analysis, protein expression, and comprehensive clinical data. The dataset has fundamentally transformed cancer research by enabling integrated analysis across molecular data types.
Relevant research paper: The Cancer Genome Atlas Pan-Cancer Analysis Project
Size: 20,000+ samples across 33 cancer types, 2.5 petabytes of data
License: Open access through dbGaP and GDC
Access the dataset: Genomic Data Commons
Ideal for: Cancer research, multi-omics integration, precision medicine, biomarker discovery, and therapeutic target identification.
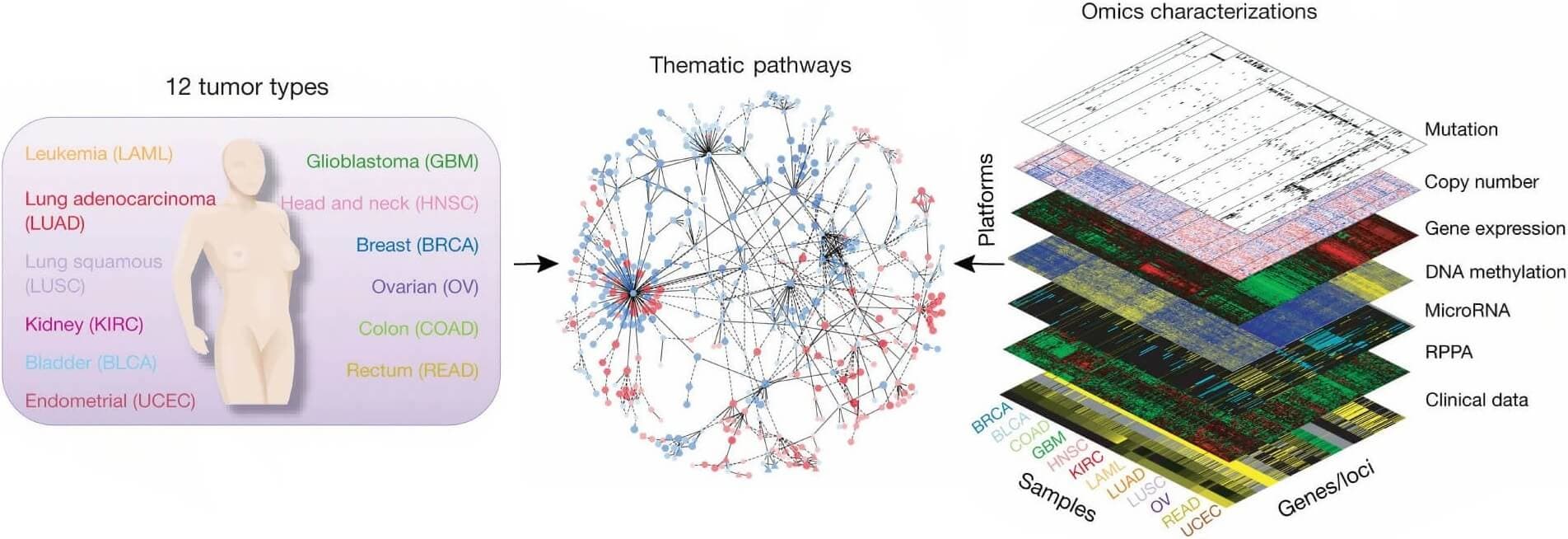
Figure 1: Integrated data set for comparing and contrasting multiple tumor types. (https://www.nature.com/articles/ng.2764)
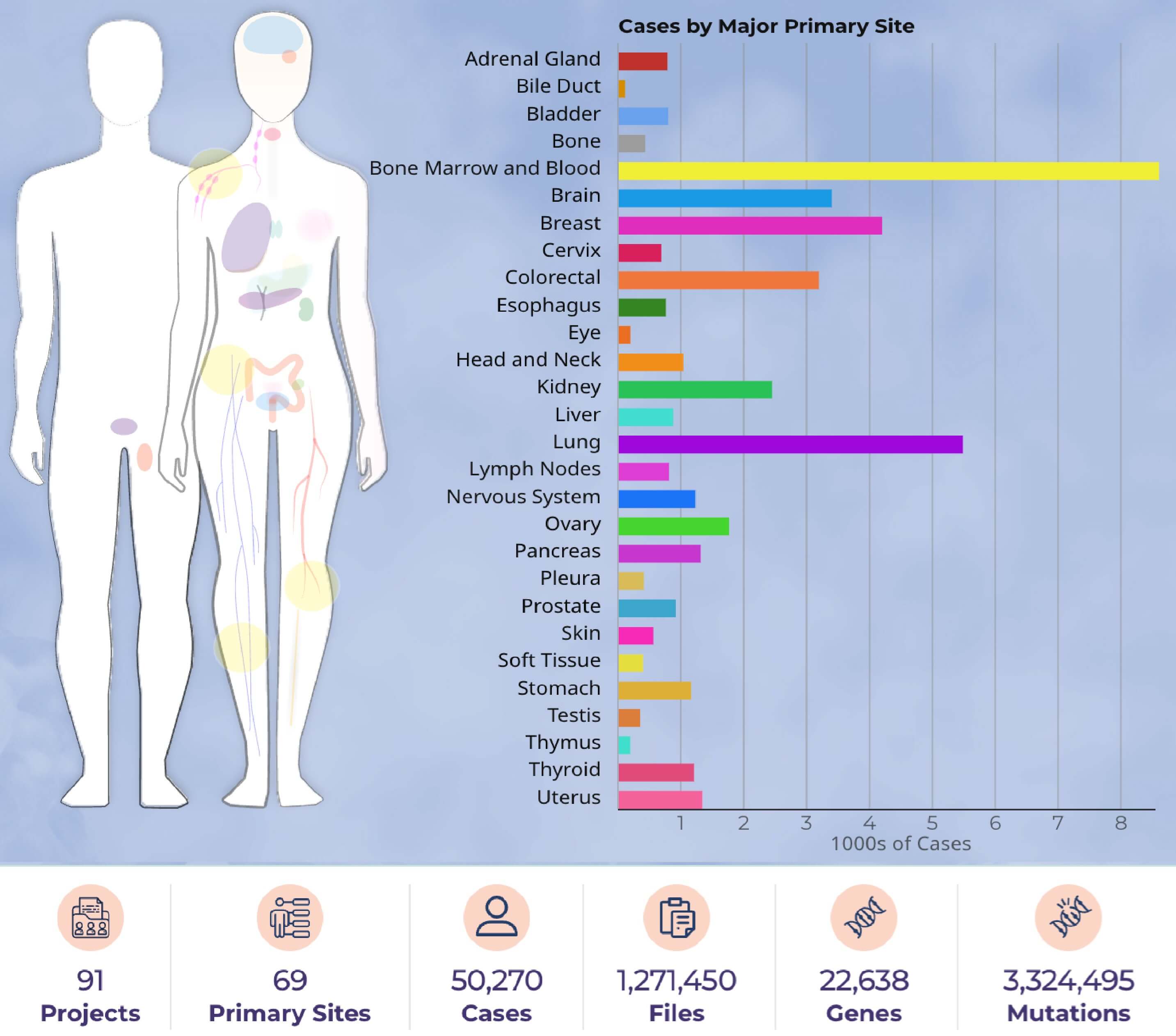
Figure 2: Cases by major primary site - Data portal summary. (https://portal.gdc.cancer.gov/)
2. CLEVR (Compositional Language and Elementary Visual Reasoning)
CLEVR is a multimodal dataset designed to evaluate a machine learning model's ability to reason about the physical world using both visual information and natural language. It is a synthetic multimodal dataset created to test AI systems' ability to perform complex reasoning about visual scenes.
CLEVR combines visual and textual modalities through rendered 3D scenes containing various objects with distinct properties like shape, size, color, and material. The textual component consists of questions posed in natural language about these scenes, challenging models to understand relationships and properties.
Relevant research paper: CLEVR: A Diagnostic Dataset for Compositional Language and Elementary Visual Reasoning
Size: 1 million images with corresponding questions
License: BSD
Access the dataset: Stanford CLEVR Dataset
Ideal for: Researchers developing visual reasoning systems, computer vision applications requiring spatial understanding, and AI models that need to process complex visual-linguistic relationships.
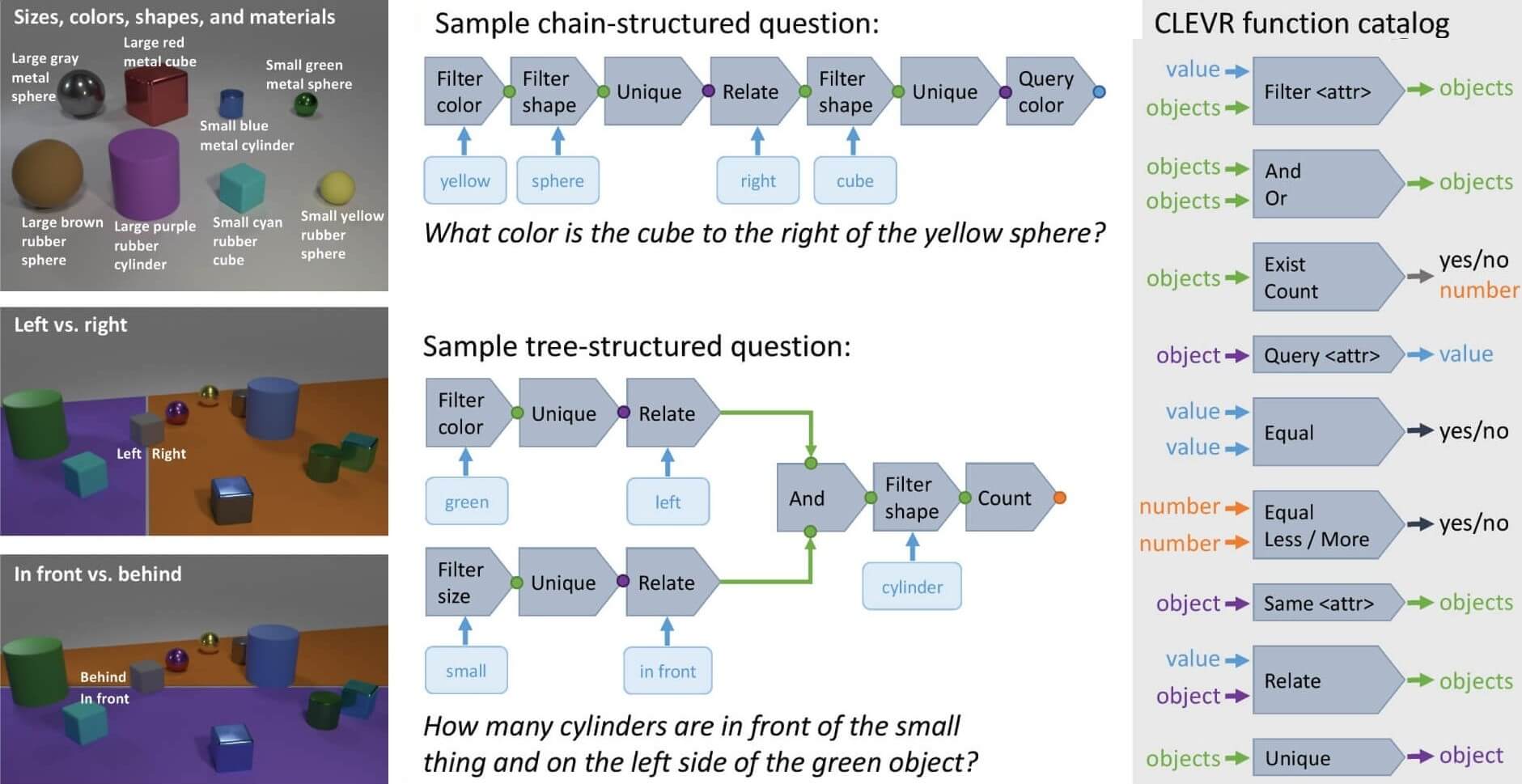
Figure 3: A field guide to the CLEVR universe. Left: Shapes, attributes, and spatial relationships. Center: Examples of questions and their associated functional programs. Right: Function catalog for building questions. (https://arxiv.org/pdf/1612.06890)
3. WIT (Wikipedia-based Image Text Dataset)
WIT primarily focuses on tasks involving the relationship between images and their textual descriptions. Some key applications are Image-Text Retrieval to retrieve images using text query, Image Captioning to generate captions for unseen images, and Multilingual Learning that can understand and connect images to text descriptions in various languages.
WIT provides a massive collection of image-text pairs extracted from Wikipedia articles across 108 languages, making it invaluable for multilingual multimodal applications.
Relevant research paper: WIT: Wikipedia-based Image Text Dataset for Multimodal Multilingual Machine Learning
Size: 37.6 million entity-rich image-text examples with 11.5 million unique images
License: Creative Commons Attribution-ShareAlike 3.0 Unported
Access the dataset: Google Research Datasets WIT
Ideal for: Multilingual AI applications, cross-modal retrieval systems, image captioning models, and researchers working on Wikipedia-scale knowledge representation.
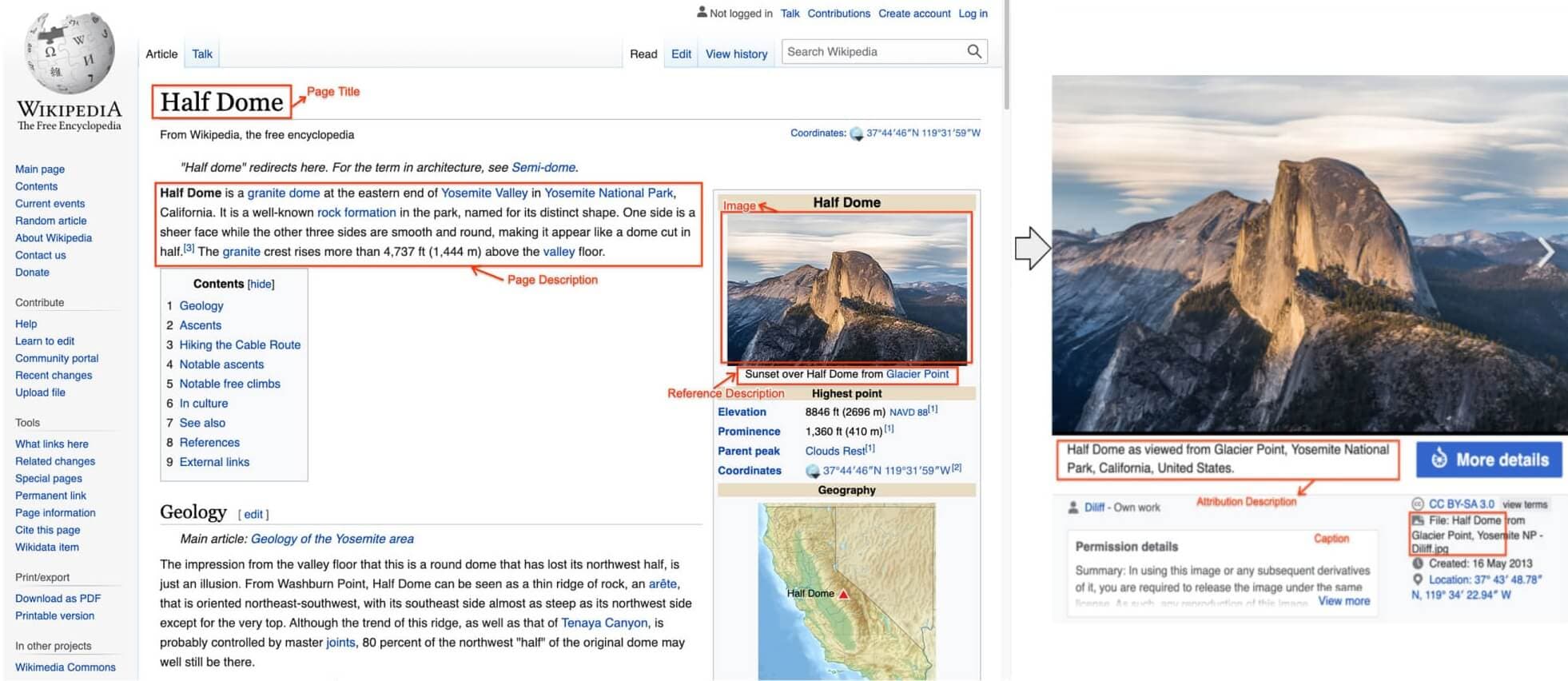
Figure 4: WIT image-text example with all text annotations. (https://arxiv.org/pdf/2103.01913)
4. LAION-5B
Developed by the LAION non-profit collective, LAION-5B is one of the largest open multimodal datasets available. It contains 5.85 billion CLIP-filtered image-text pairs extracted from the web, with 2.32 billion English pairs and the rest spanning multiple languages.
LAION-5B democratizes access to large-scale data used for training foundation models like CLIP and Stable Diffusion, providing researchers with web-scale multimodal data.
Relevant research paper: LAION-5B: An open large-scale dataset for training next generation image-text models
Size: 5.85 billion image-text pairs
License: Various (depends on source)
Access the dataset: LAION Dataset
Ideal for: Training large-scale foundation models, text-to-image generation, multimodal representation learning, and researchers requiring massive datasets for model development.

Figure 5: LAION-5B examples. Sample images from a nearest neighbor search in LAION-5B using CLIP embeddings. The image and caption (C) are the first results for the query (Q). (https://arxiv.org/pdf/2210.08402)
5. MS COCO (Microsoft Common Objects in Context)
MS COCO provides comprehensive annotations for object detection, segmentation, and captioning tasks. The dataset contains images with multiple objects in natural contexts, accompanied by detailed annotations and captions.
Relevant research paper: Microsoft COCO: Common Objects in Context
Size: 330,000 images with over 2.5 million labeled instances
License: Creative Commons Attribution 4.0
Access the dataset: COCO Dataset
Ideal for: Object detection research, image segmentation, caption generation, and computer vision benchmarking.
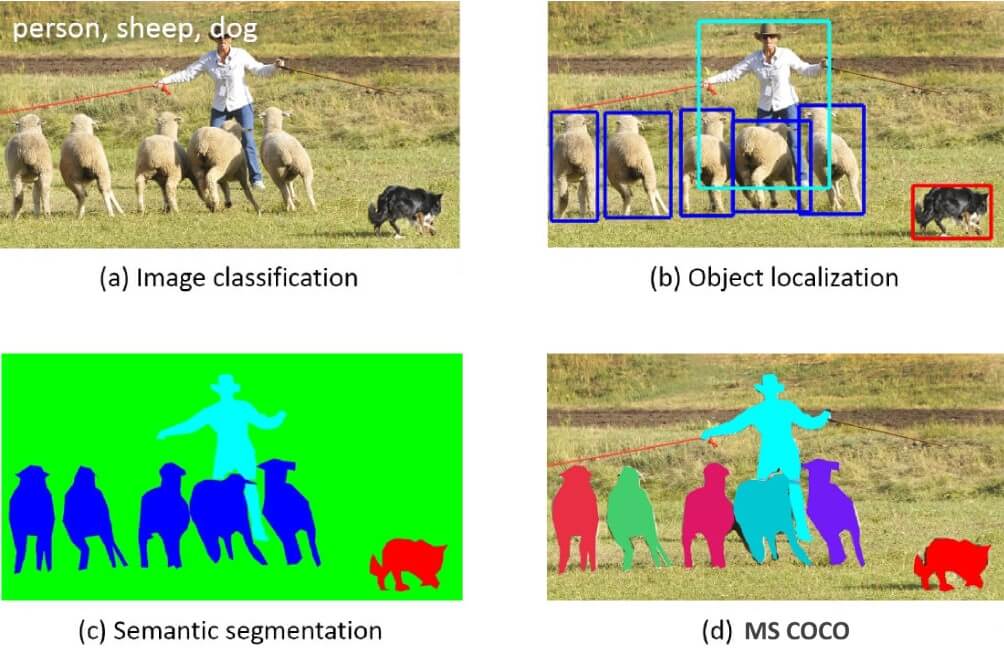
Figure 6: Previous object recognition datasets focus on (a), (b) and (c), MS COCO focuses on (d) segmenting individual object instances, by introducing a large, richly-annotated dataset comprised of images depicting complex everyday scenes of common objects in their natural context. (https://arxiv.org/pdf/1405.0312)
6. VQA (Visual Question Answering)
VQA combines visual and textual information to enable AI systems to answer questions about images. The dataset requires models to understand both visual content and natural language questions.
Relevant research paper: VQA: Visual Question Answering
Size: 265,016 images with over 614,000 questions
License: Creative Commons Attribution 4.0
Access the dataset: VQA Dataset
Ideal for: Visual question answering research, multimodal reasoning systems, and educational AI applications.

Figure 7: Examples of free-form, open-ended questions collected for images via Amazon Mechanical Turk. Note that commonsense knowledge is needed along with a visual understanding of the scene to answer many questions. (https://arxiv.org/pdf/1505.00468)
7. MIMIC-IV
MIMIC-IV provides multimodal medical data including clinical notes, lab results, vital signs, medications, and procedures from intensive care unit patients. The dataset enables comprehensive healthcare AI research by combining structured and unstructured clinical data.
MIMIC-IV represents one of the most complete multimodal clinical datasets available, integrating time-series physiological data with rich textual clinical documentation and structured medical records.
Relevant research paper: MIMIC-IV, a freely accessible electronic health record dataset
Size: 400,000+ hospital admissions with comprehensive clinical data
License: PhysioNet Credentialed Health Data License
Access the dataset: MIMIC-IV PhysioNet
Ideal for: Healthcare AI research, clinical decision support systems, medical predictive modeling, and natural language processing in healthcare.
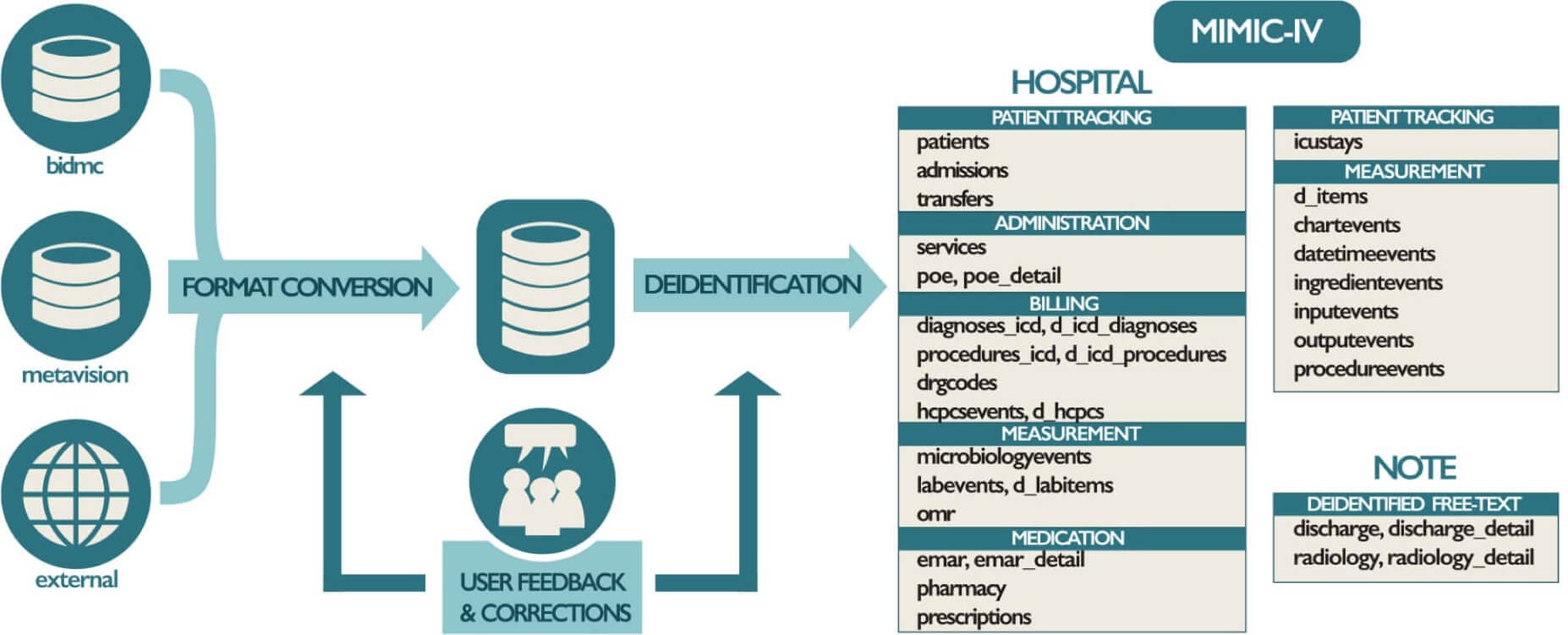
Figure 8: An overview of the development process for MIMIC-IV. (https://www.nature.com/articles/s41597-022-01899-x.pdf)
8. UK Biobank
UK Biobank provides genomic data, medical imaging (MRI, retinal), proteomics, clinical records, and lifestyle data from 500,000+ participants with longitudinal follow-up. This massive prospective cohort study combines genetic, environmental, and lifestyle factors with detailed health outcomes.
The dataset includes whole genome sequencing, multi-organ MRI imaging, retinal photography, and comprehensive phenotyping data, making it invaluable for understanding gene-environment interactions in health and disease.
Relevant research paper: The UK Biobank resource with deep phenotyping and genomic data
Size: 500,000+ participants with genomic, imaging, and clinical data
License: Approved researcher access
Access the dataset: UK Biobank
Ideal for: Population genomics, precision medicine, epidemiological studies, and gene-environment interaction research.
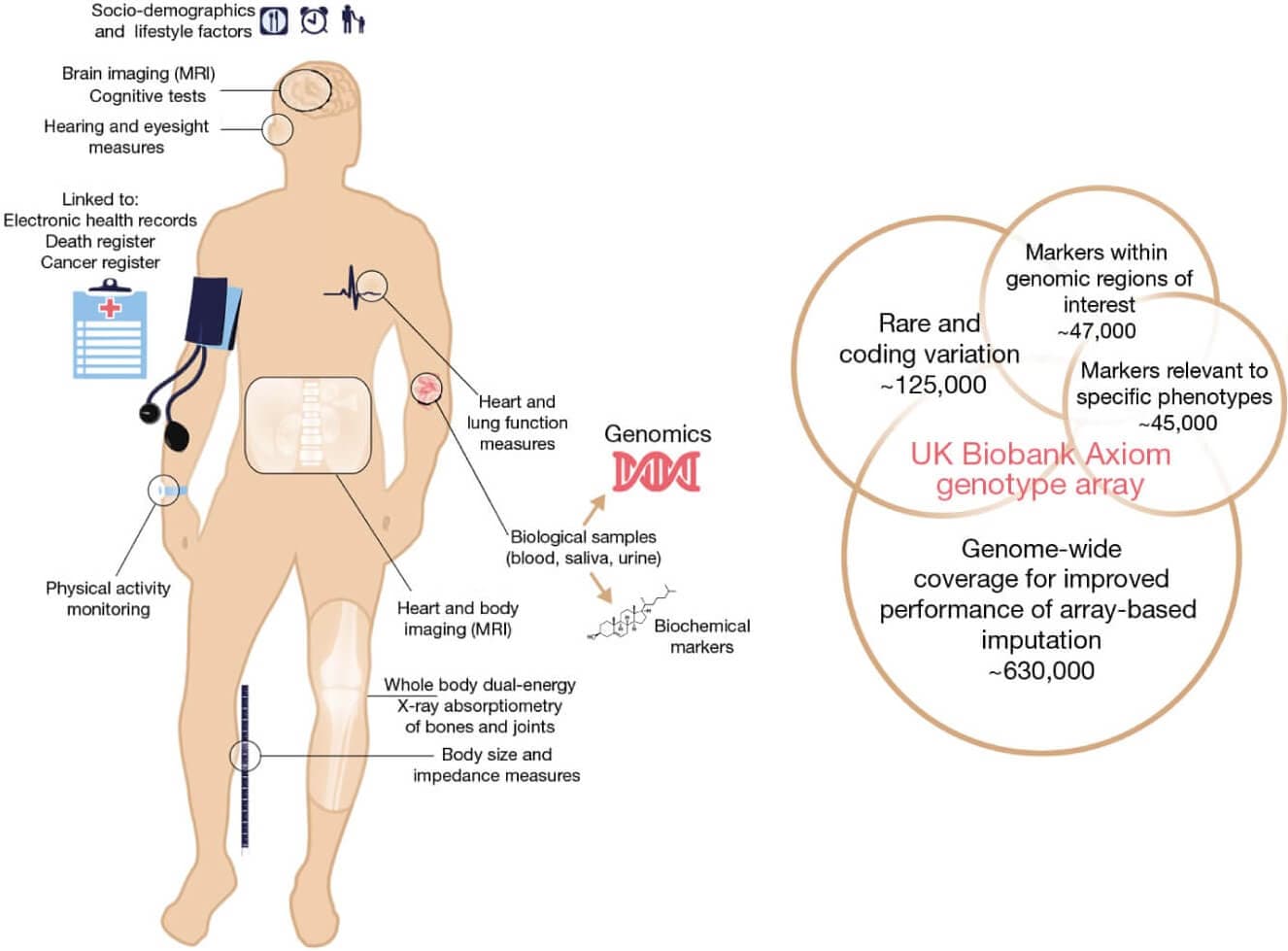
Figure 9: Summary of the UK Biobank resource and genotyping array content. (http://nature.com/articles/s41586-018-0579-z)
9. Human Cell Atlas (HCA)
The Human Cell Atlas integrates single-cell RNA sequencing, spatial transcriptomics, proteomics, and imaging data to create comprehensive maps of all human cells. This international collaboration aims to map every cell type in the human body across development, health, and disease.
HCA combines cutting-edge single-cell technologies with spatial information to understand cellular diversity, tissue organization, and developmental processes at unprecedented resolution.
Relevant research paper: The Human Cell Atlas
Size: Millions of cells across multiple tissues and developmental stages
License: Open access with data use agreements
Access the dataset: Human Cell Atlas Data Portal
Ideal for: Single-cell biology, developmental biology, disease research, and understanding cellular diversity in human tissues.
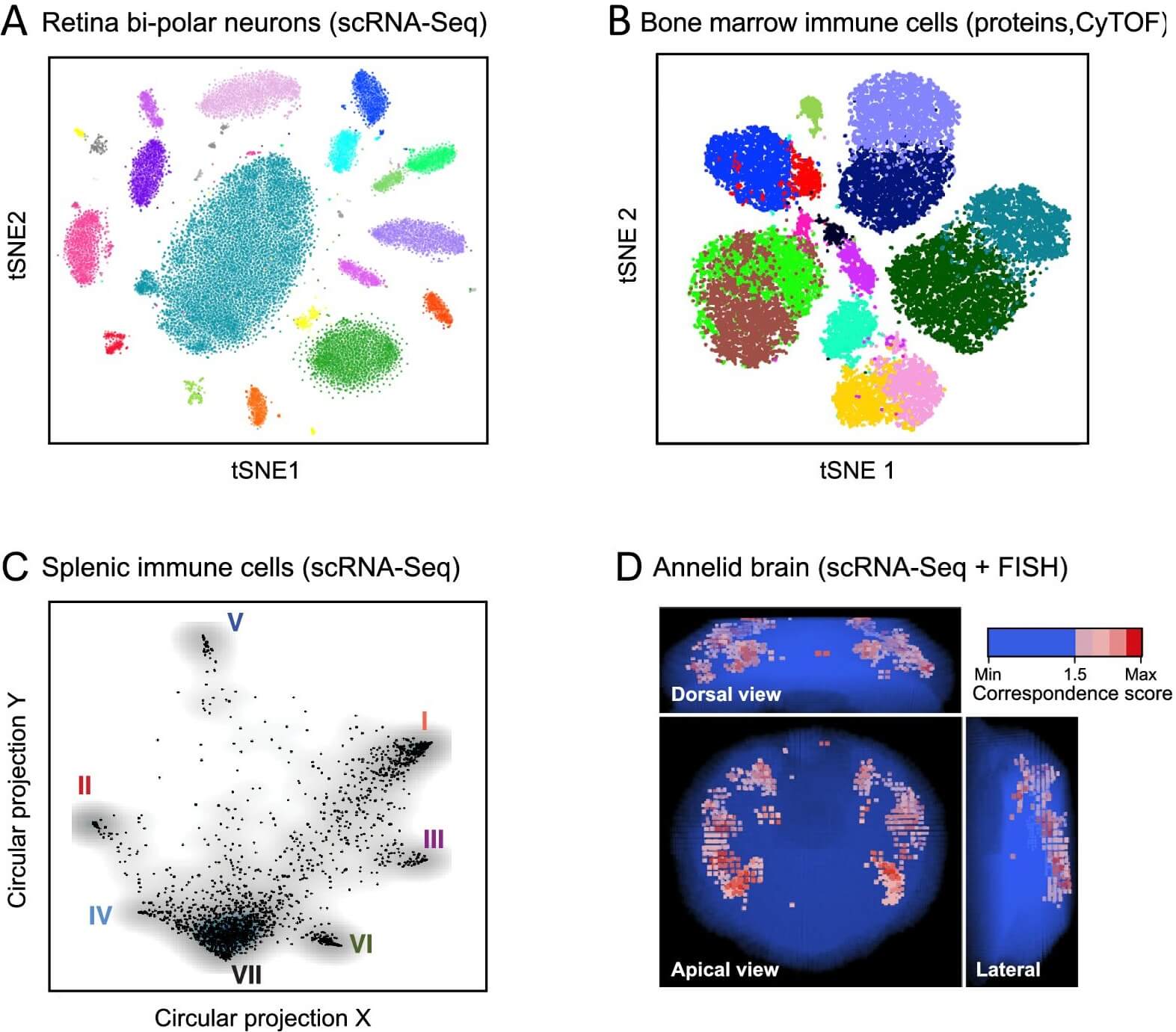
Figure 10: Anatomy: cell types and tissue structure. (https://elifesciences.org/articles/27041)
10. TCIA (The Cancer Imaging Archive)
The compiled datasets encompass a broad spectrum of data modalities, such as radiology images (CT, MRI, PET), pathology slides, genomic data, and clinical records. This multimodal nature enables the integration of different data types to capture the intricacies of cancer.
TCIA provides medical imaging data paired with clinical information for cancer research and diagnosis development.
Relevant research paper: The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository
Size: 30+ million images across multiple cancer types
License: Various (most are open access)
Access the dataset: The Cancer Imaging Archive
Ideal for: Medical AI research, cancer diagnosis systems, radiological analysis, and imaging-genomics integration.
11. MLOmics
MLOmics is an open cancer multi-omics database specifically designed for machine learning applications. The dataset contains genomic, transcriptomic, proteomic, and clinical data from cancer patients, with extensive preprocessing and baseline models for immediate use in ML workflows.
MLOmics addresses the gap between raw omics data and machine learning-ready datasets by providing standardized, well-annotated multimodal cancer data with comprehensive baseline evaluations.
Relevant research paper: MLOmics: Cancer Multi-Omics Database for Machine Learning
Size: 8,314 patient samples covering 32 cancer types with four omics types
License: Open access
Access the dataset: MLOmics Database
Ideal for: Cancer machine learning research, multi-omics integration, biomarker discovery, and developing AI models for precision oncology.
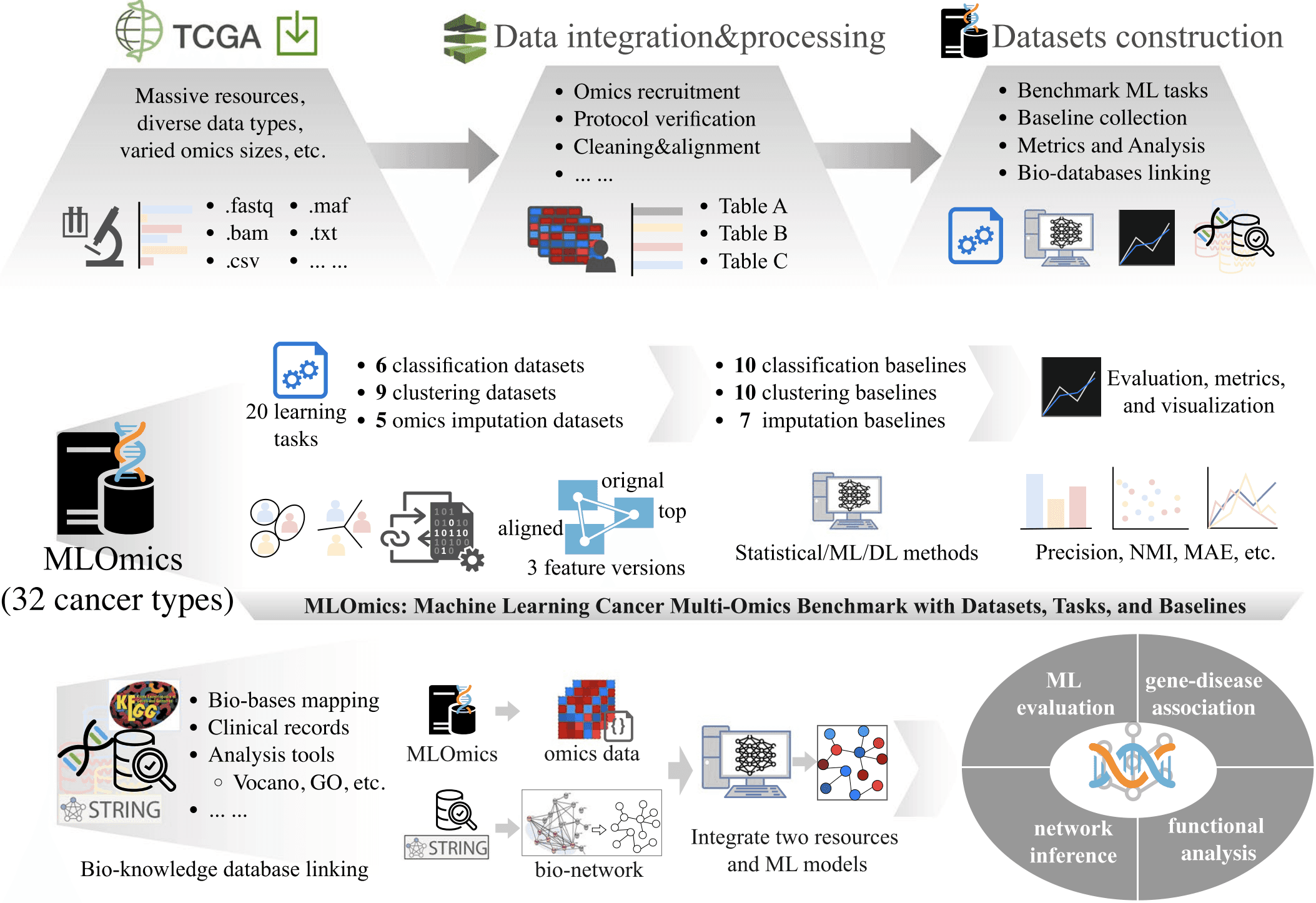
Figure 11: Schematic workflow of creating the MLOmics. (https://www.nature.com/articles/s41597-025-05235-x.pdf)
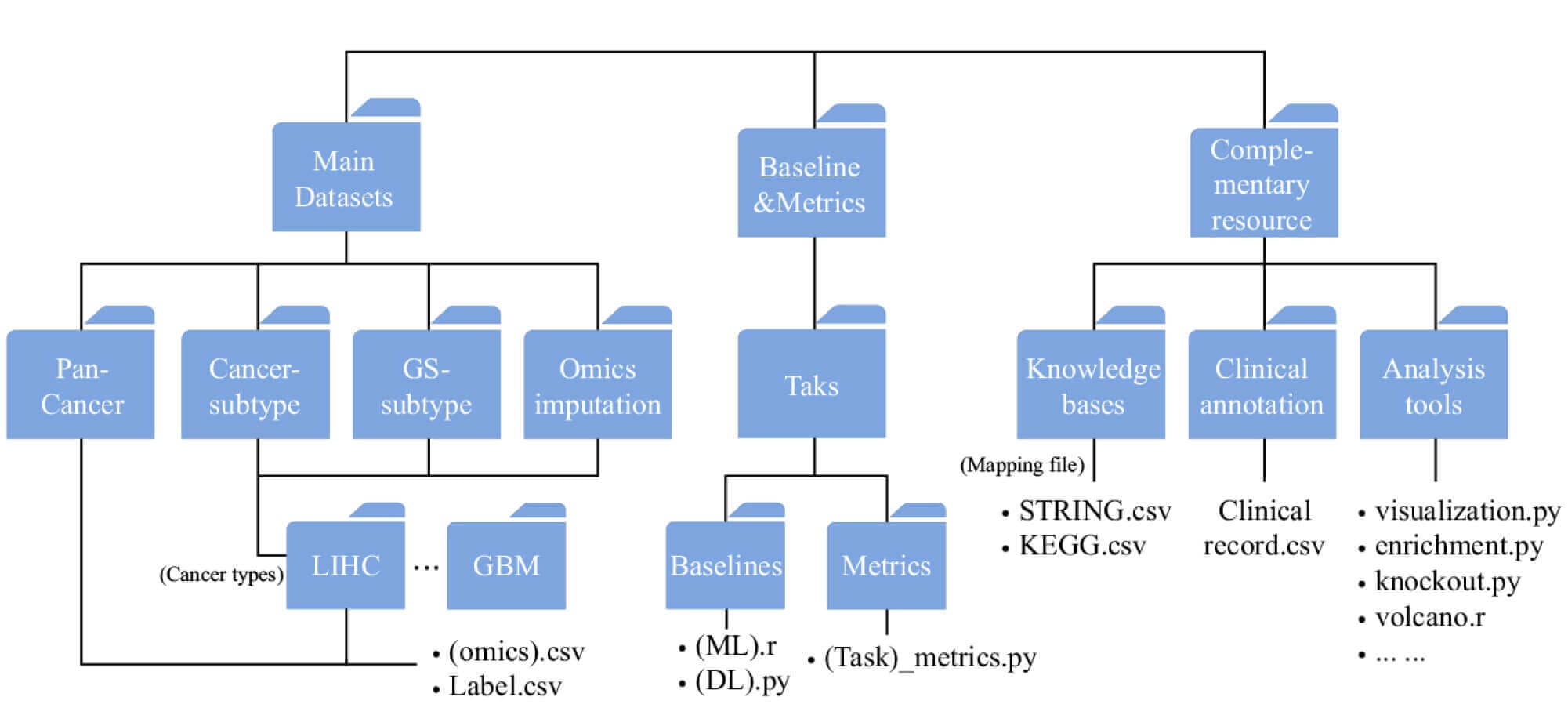
Figure 12: Schematic of MLOmics resources structure. (https://www.nature.com/articles/s41597-025-05235-x.pdf)
12. Kinetics-700
Kinetics-700 contains human action videos across 700 action categories, providing rich temporal visual data for action recognition and video understanding research.
Relevant research paper: The Kinetics Human Action Video Dataset
Size: 650,000 video clips
License: YouTube terms of service
Access the dataset: DeepMind Kinetics
Ideal for: Action recognition research, video classification, and temporal modeling in computer vision.

Figure 13: Example classes from the Kinetics dataset. Note that in some cases a single image is not enough for recognizing the action or distinguishing classes. (https://arxiv.org/pdf/1705.06950)
13. Flickr30k
Flickr30k contains images from Flickr paired with human-written captions, providing high-quality image-text associations for multimodal learning.
Relevant research paper: From image descriptions to visual denotations: New similarity metrics for semantic inference over event descriptions
Size: 31,000 images with 158,000 captions
License: Academic research
Access the dataset: Flickr30k Dataset
Ideal for: Image-text retrieval, caption generation, and visual-semantic understanding research.

Figure 14: Two images from the data set and their five captions. (https://aclanthology.org/Q14-1006.pdf)
14. ENCODE
ENCODE provides comprehensive data on gene regulation through ChIP-seq, RNA-seq, chromatin accessibility assays, and histone modification mapping across cell types and conditions. The project aims to identify all functional elements in the human genome.
ENCODE integrates multiple genomic assays to create comprehensive maps of gene regulatory networks, combining transcriptional, epigenetic, and chromatin accessibility data.
Relevant research paper: An integrated encyclopedia of DNA elements in the human genome
Size: 1,600+ experiments across multiple cell types and conditions
License: Open access
Access the dataset: ENCODE Portal
Ideal for: Gene regulation research, epigenomics studies, regulatory network analysis, and understanding functional genomics.
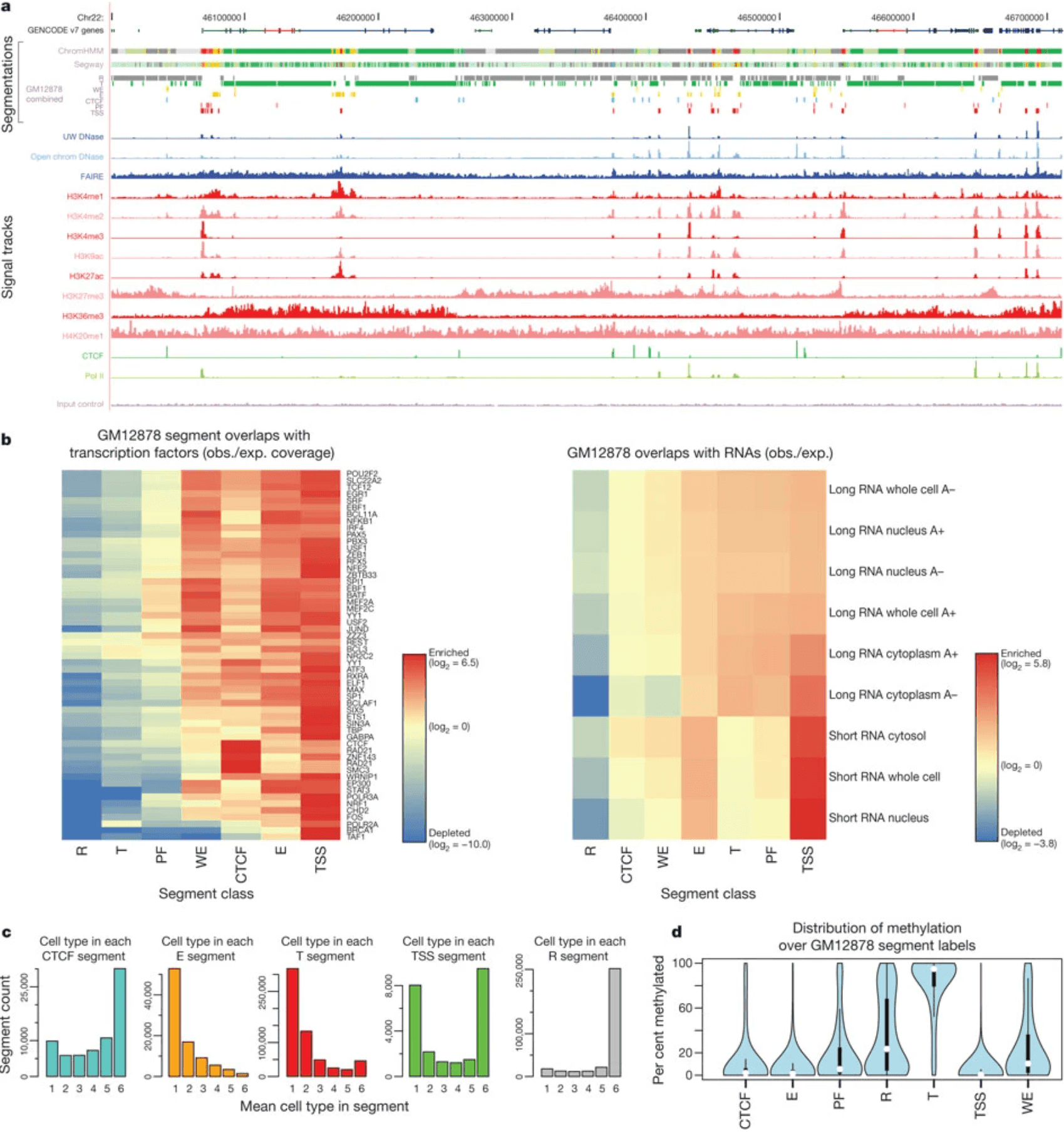
Figure 15: Integration of ENCODE data by genome-wide segmentation. (https://www.nature.com/articles/nature11247)
15. Allen Brain Atlas
The Allen Brain Atlas combines gene expression data, neuroimaging, connectivity mapping, and cell type characterization across brain regions and developmental stages. This comprehensive resource provides multimodal data for understanding brain structure and function.
The atlas integrates anatomical, molecular, and functional data to create detailed maps of brain organization across species and developmental time points.
Relevant research paper: Genome-wide atlas of gene expression in the adult mouse brain
Size: Multiple brain regions across development and disease states
License: Open access
Access the dataset: Allen Brain Atlas Portal
Ideal for: Neuroscience research, brain development studies, neurological disease research, and systems neuroscience.
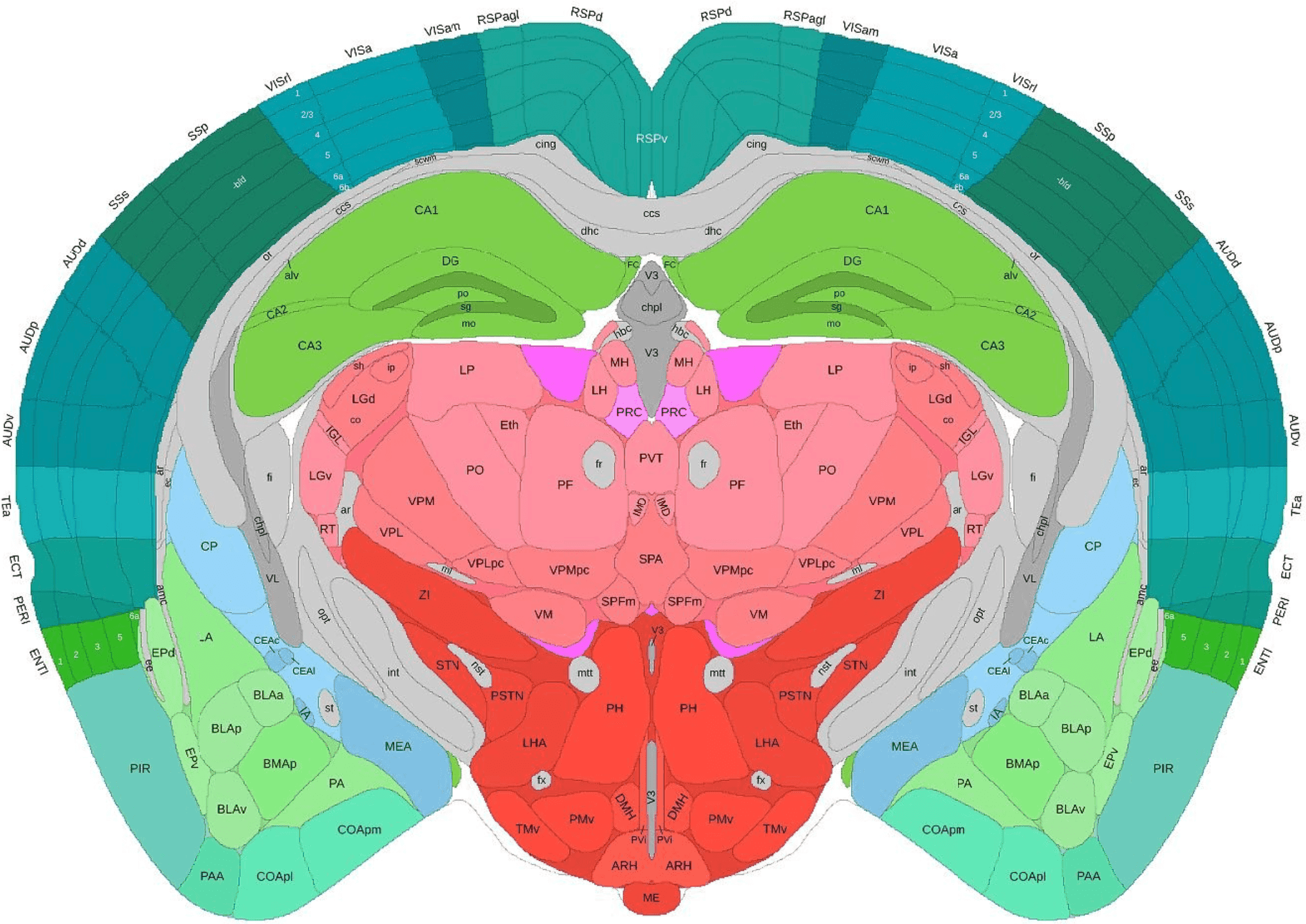
Figure 16: Adult mouse 3D coronal.
Are there free multimodal datasets?
Yes, there are numerous free multimodal datasets available for research and development purposes. Many prominent datasets in this guide are freely accessible, including:
TCGA - Available through open access portals
CLEVR - Released under BSD license
Human Cell Atlas - Open access with data use agreements
Free datasets may have limitations such as restricted commercial use, academic-only licensing, or requirements for attribution. Researchers can find free datasets through academic institutions, government organizations, and open-source communities.
How can you find multimodal datasets?
You can find multimodal datasets through specialized repositories, academic institutions, and research organizations.
Academic repositories: Papers with Code, Google Dataset Search, and university research labs
Government sources: NIH data repositories, cancer genomics portals, and national research initiatives
Industry platforms: Kaggle, Hugging Face, and company research divisions
Specialized collections: Medical imaging archives, genomics databases, and domain-specific repositories
How can you analyze multimodal datasets?
To analyze multimodal datasets, you need specialized platforms and tools capable of handling heterogeneous data types simultaneously. Traditional database systems struggle with the complexity and scale of multimodal data, particularly in life sciences where datasets may include genomics, imaging, and clinical data.
Modern multimodal analysis requires platforms that can efficiently store, index, and query across different data modalities. These systems must handle the temporal synchronization between modalities and provide unified access to disparate data types.
TileDB provides a comprehensive platform for multimodal data analysis through its multi-dimensional array architecture. The platform enables organizations to store, organize, and analyze complex multimodal datasets efficiently, supporting everything from medical imaging combined with genomic data to single-cell multiomics with spatial information.
For organizations looking to implement multimodal data analysis capabilities, TileDB offers enterprise-grade solutions that scale from research projects to production deployments. Contact our sales team to learn how TileDB can accelerate your multimodal data initiatives.
Which industries use multimodal datasets?
Multimodal datasets are transforming industries by enabling more comprehensive analysis and decision-making across sectors.
Academic research: Universities combine imaging, genomics, and clinical data for breakthrough medical discoveries
Biotechnology: Single-cell multiomics integrates genomics, proteomics, and spatial data for understanding cellular biology
Diagnostics: Medical device manufacturers combine imaging data with AI models for automated diagnosis
Healthcare: Multimodal datasets can combine medical imaging, genomic data, patient records, and clinical notes for improved diagnosis and treatment planning. This multimodal approach enables the integration of different data types to capture the intricacies of cancer and other complex medical conditions.
Healthcare systems: Hospitals integrate electronic health records, medical imaging, and laboratory data for clinical decision support
Pharmaceutical: Drug discovery combines molecular data, imaging, genomics, and clinical trial data for precision medicine approaches
Additional industries adopting multimodal approaches include agriculture (combining satellite imagery with genomics data for crop improvement), environmental science (integrating sensor data with imaging for climate research), and materials science (combining microscopy with molecular data for material discovery).
Multimodal datasets enable these industries to build AI systems that mirror human-like understanding by processing information from multiple sources simultaneously. Organizations implementing multimodal approaches often see significant improvements in accuracy and insights compared to single-modality systems.
Ready to harness the power of multimodal data for your organization? TileDB provides the infrastructure and expertise needed to successfully implement multimodal data solutions across life sciences and beyond. Contact our team to discuss your specific requirements and learn how we can accelerate your multimodal AI initiatives.
Meet the authors


